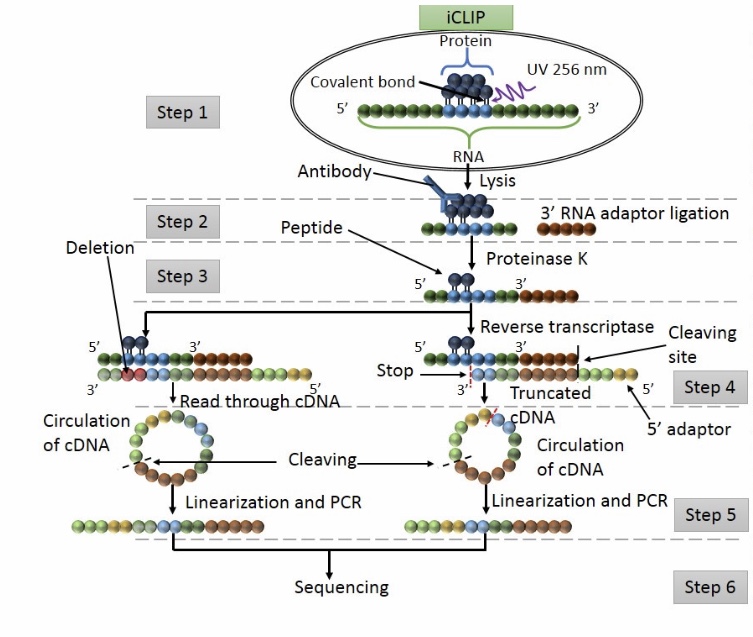
Enhanced Crosslinking Immunoprecipitation (eCLIP) Method for Efficient Identification of Protein-bound RNA in Mouse Testis | Protocol
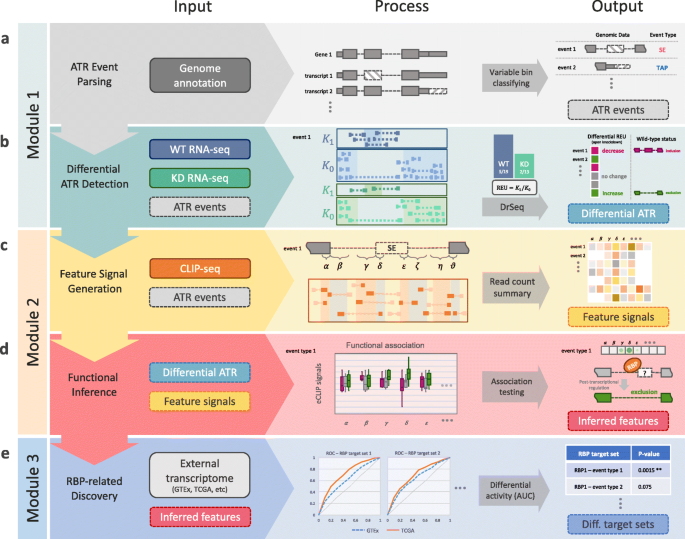
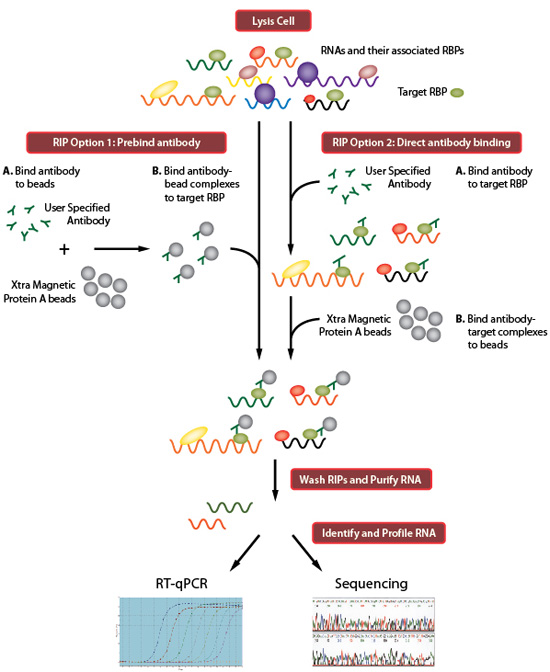
SURF: integrative analysis of a compendium of RNA-seq and CLIP-seq datasets highlights complex governing of alternative transcriptional regulation by RNA-binding proteins | Genome Biology | Full Text

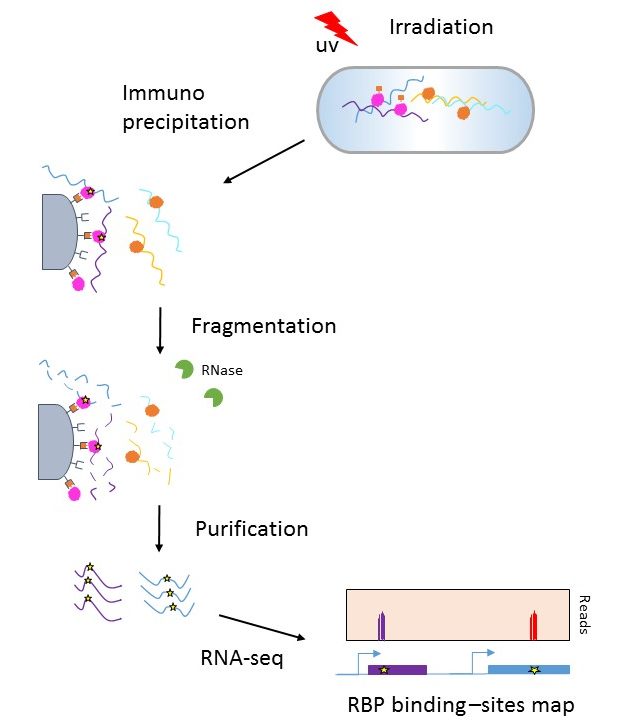
Global RNA recognition patterns of post‐transcriptional regulators Hfq and CsrA revealed by UV crosslinking in vivo | The EMBO Journal

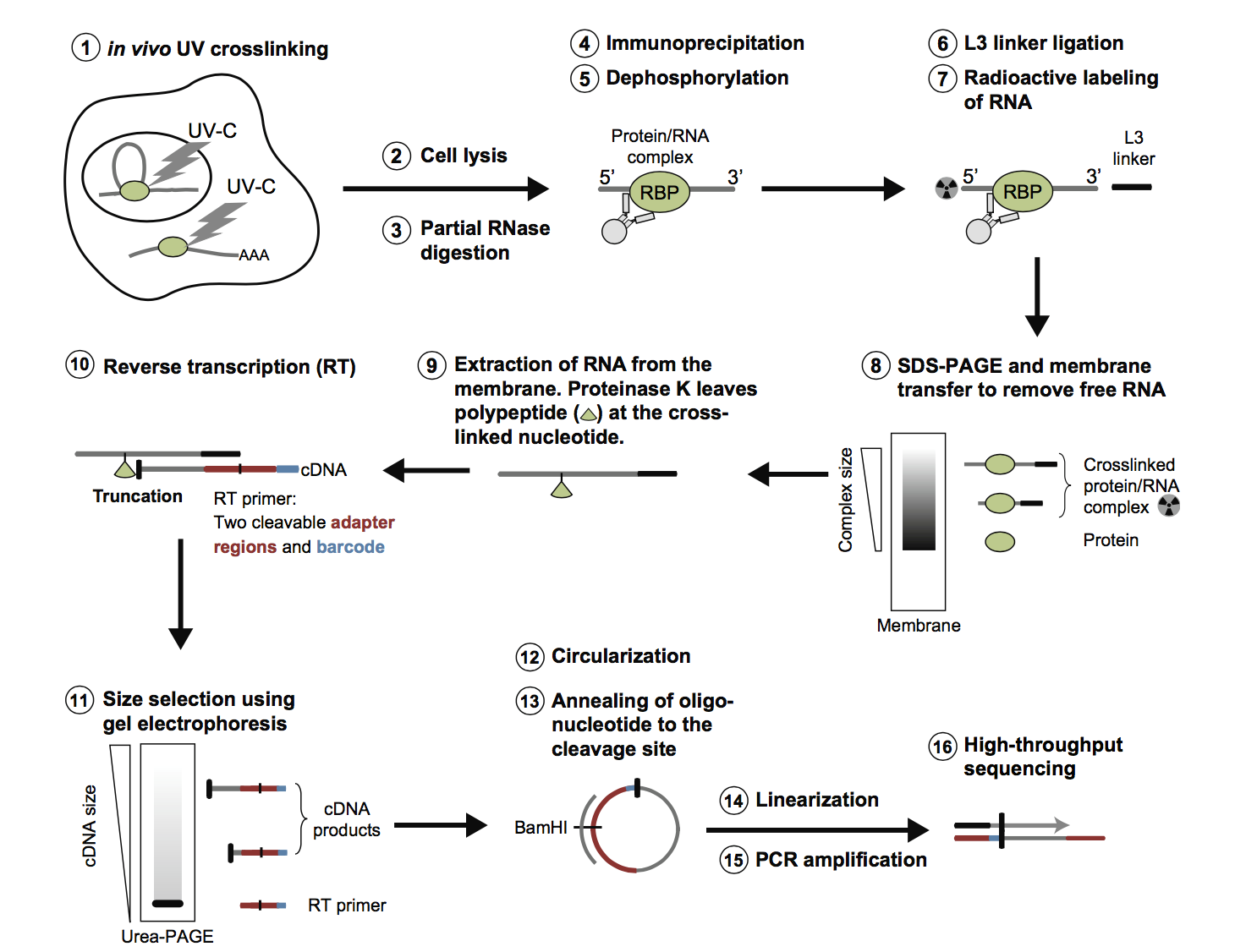
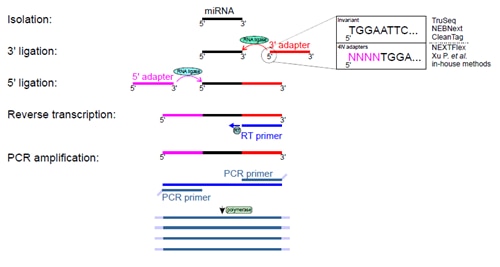
PAR-CLIP and Streamlined Small RNA cDNA Library Preparation Protocol for the Identification of RNA Binding Protein Target Sites | RNA-Seq Blog

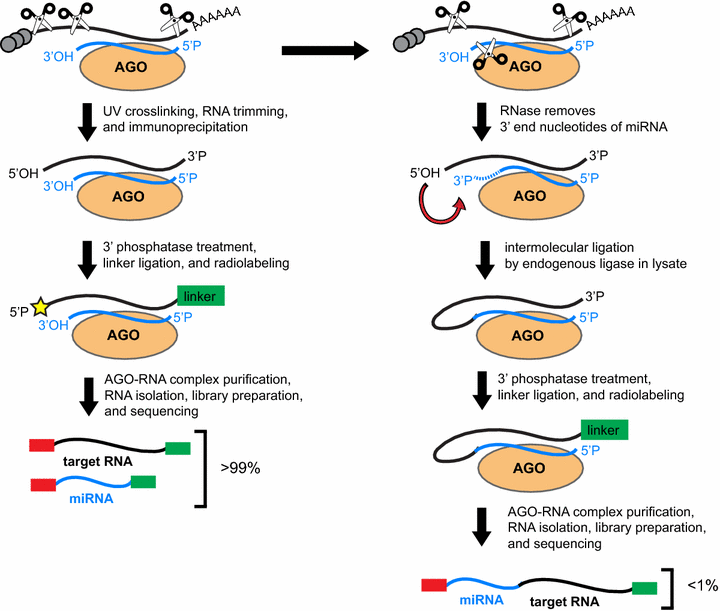
Pipeline for extracting MIWI CLIP-Seq features (see Section 2). The RNA... | Download Scientific Diagram

easyCLIP Quantifies RNA-Protein Interactions and Characterizes Recurrent PCBP1 Mutations in Cancer | bioRxiv
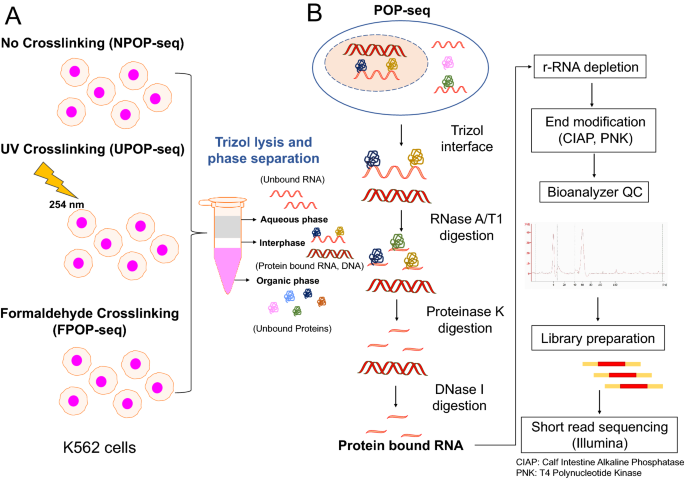
Transcriptome-wide high-throughput mapping of protein–RNA occupancy profiles using POP-seq | Scientific Reports